Note
Go to the end to download the full example code.
Field data inversion (“Koenigsee”)#
This minimalistic example shows how to use the Refraction Manager to invert a field data set. Here, we consider the Koenigsee data set, which represents classical refraction seismics data set with slightly heterogeneous overburden and some high-velocity bedrock. The data file can be found in the pyGIMLi example data repository.
# We import pyGIMLi and the traveltime module.
import matplotlib.pyplot as plt
import pygimli as pg
import pygimli.physics.traveltime as tt
The helper function pg.getExampleData downloads the data set to a temporary location and loads it. Printing the data reveals that there are 714 data points using 63 sensors (shots and geophones) with the data columns s (shot), g (geophone), and t (traveltime). By default, there is also a validity flag.
[::::::::::::::::::::::::::::::::::::: 83% ::::::::::::::::::::::: ] 8193 of 9844 complete
[:::::::::::::::::::::::::::::::::::: 100% ::::::::::::::::::::::::::::::::::::] 9844 of 9844 complete
md5: 641890bb17cb2bdf052cbc348669dfd0
Data: Sensors: 63 data: 714, nonzero entries: ['g', 's', 't', 'valid']
Let’s have a look at the data in the form of traveltime curves.
fig, ax = plt.subplots()
lines = tt.drawFirstPicks(ax, data)
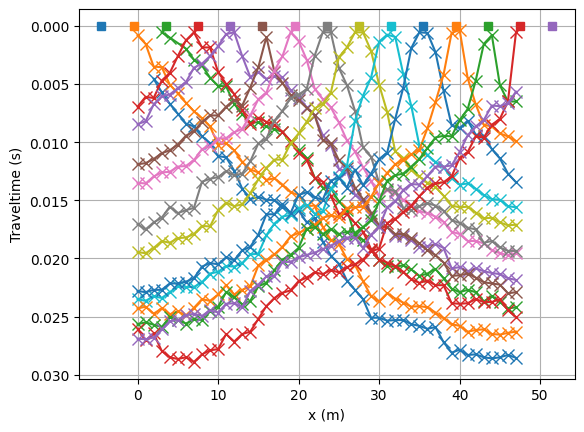
We initialize the refraction manager.
mgr = tt.TravelTimeManager(data)
# Alternatively, one can plot a matrix plot of apparent velocities which is the
# more general function also making sense for crosshole data.
ax, cbar = mgr.showData()
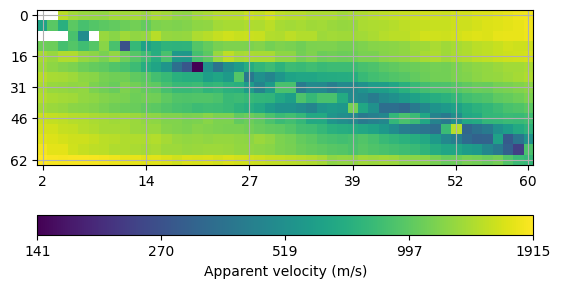
Finally, we call the invert method and plot the result.The mesh is created based on the sensor positions on-the-fly.
mgr.invert(secNodes=3, paraMaxCellSize=5.0,
zWeight=0.2, vTop=500, vBottom=5000, verbose=1)
--------------------------------------------------------------------------------
--------------------------------------------------------------------------------
inv.iter 1 ... --------------------------------------------------------------------------------
inv.iter 2 ... --------------------------------------------------------------------------------
inv.iter 3 ... --------------------------------------------------------------------------------
inv.iter 4 ... --------------------------------------------------------------------------------
inv.iter 5 ... --------------------------------------------------------------------------------
inv.iter 6 ... --------------------------------------------------------------------------------
inv.iter 7 ... --------------------------------------------------------------------------------
inv.iter 8 ... --------------------------------------------------------------------------------
inv.iter 9 ... --------------------------------------------------------------------------------
inv.iter 10 ... --------------------------------------------------------------------------------
inv.iter 11 ... --------------------------------------------------------------------------------
inv.iter 12 ... --------------------------------------------------------------------------------
inv.iter 13 ... ################################################################################
# Abort criterion reached: dPhi = 1.98 (< 2.0%) #
################################################################################
1090 [898.9879603451514,...,2649.5722103614794]
Look at the fit between measured (crosses) and modelled (lines) traveltimes.
mgr.showFit(firstPicks=True)
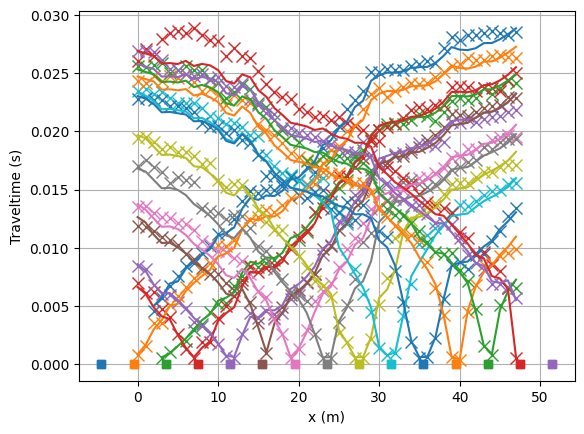
You can plot only the model and customize with a bunch of keywords
ax, cbar = mgr.showResult(logScale=False, cMin=500, cMax=3000, cMap="plasma_r",
coverage=mgr.standardizedCoverage())
rays = mgr.drawRayPaths(ax=ax, color="k", lw=0.3, alpha=0.5)
# mgr.coverage() yields the ray coverage in m and standardizedCoverage as 0/1
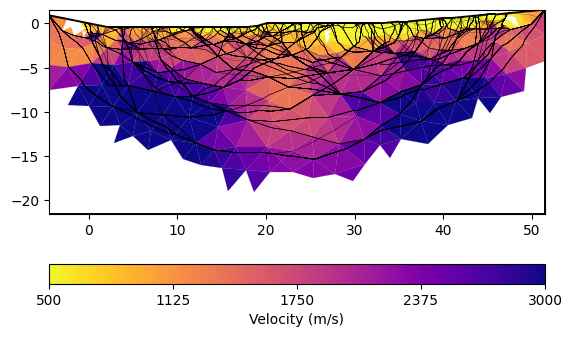
You can play around with the gradient starting model (vTop and vBottom arguments) and the regularization strength lam and customize the mesh.
Total running time of the script: (0 minutes 34.205 seconds)